The threat of antibiotic resistance continues to grow. A world without antibiotics is one where 3 million common procedures per year, such as caesarean sections and hip replacements become life-threatening, and cancer patients are put at further risk.
Antibiotic-resistant bloodstream infections rose by an estimated 35% between 2013 and 2017, and we are seeing an increasing number of infections that are not responding to antibiotics of last resort. It is clear that innovative new approaches are needed urgently.
New funding for the most advanced data collection
Public Health England has been awarded £5.1 million by the Department of Health and Social Care to stimulate a number of initiatives, including the creation of the most advanced dataset globally on antimicrobial resistance (AMR). These funds will enable us to create a virtual ‘open access’ centre, which will gather real-time patient data on resistant infections, helping clinicians understand when to use and preserve antibiotics in their treatment. This will provide a unique and tailored approach to the huge challenge AMR poses.
Big Data to tackle big questions
There a range of different data sets available that help us interrogate antibiotic prescribing and AMR.
AMR data from PHE microbiology systems, hospital episode statistics, patient level prescribing data, and ONS mortality data are all important tools for government and researchers to monitor and probe the impact of AMR in this country.
What is missing is a way to effectively integrate all this information in a meaningful and secure manner. The new funds will allow PHE to address these challenges by creating a new dataset that is the most advanced globally. The funds will also ensure that the UK leads the development of Big Data methodologies, and essentially enhance our understanding of AMR.
The wealth of data will be used to create an open-access simulated dataset that can be used by PHE, the NHS and external researchers to comprehensively understand the drivers of antibiotic prescribing, and the impact of prescribing decisions on AMR and long-term health outcomes. The project aims to have a simulated dataset available in early 2021.
These insights will also be invaluable for policymakers in shaping guidelines that limit antibiotic consumption to the safest level.
Predicting patient responses to improve and tailor treatment
The concept of precision medicine has been defined as a set of prevention and treatment strategies that take individual variability into account. It is at a basic level tailored medicine and treatment for individuals, taking into account that we are all different with different needs.
While these approaches are often thought of in the context of non-communicable diseases such as cardiovascular disease and cancer, there is also scope to use them within the world of infectious diseases. By understanding which patients develop infections that require hospitalisation or are antibiotic resistant we can target effective treatment to those at most risk.
Drilling down further into the dataset to patient-level data will ultimately enable healthcare professionals to make decisions based on patient history. A complete picture of previous infections, antibiotics prescribed and the outcomes of those interventions, coupled with new insights into the risk factors for acquiring an antibiotic resistant infection, will enable us to predict individual patient responses and give people effective and personalised treatment.
Reducing healthcare-associated infections (HCAIs)
Reducing antibiotic prescribing is just one part of the challenge surrounding AMR – reducing infections in the first place and developing and evaluating new therapeutics are also vital.
Core infection control measures such as hand hygiene, isolation and environmental cleaning have driven down Clostridium difficile and Methicillin-resistant Staphylococcus aureus (MRSA) infection rates in hospital. MRSA blood stream infections now impact 1.5 people per 100,000 of the population compared to its peak of 15.4 per 100,000 in 2004.
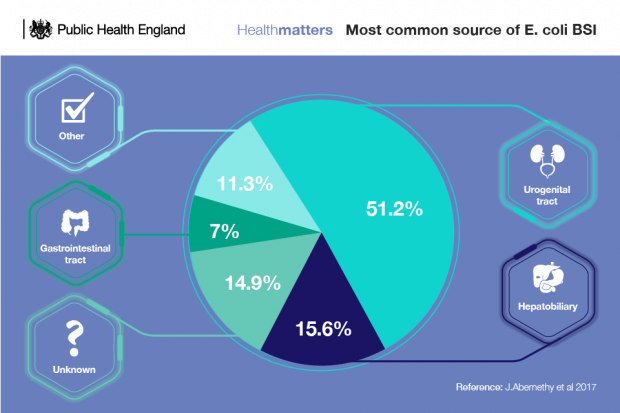
However, despite robust infection prevention control practices and relentless focus on driving down HCAIs across the healthcare system, interventions are not having a large enough impact on infections such as E. coli bloodstream infections.
The role of the hospital environment in the transmission of HCAIs is still not fully understood. In order to address this, PHE will establish a flexible modular facility that will simulate a 4-bed ward (a facility that looks like a normal ward, but exists for research purposes), isolation room with lobby and assisted shower room, among other things. To do this, we will draw on our experience of designing, delivering and operating similar modular facilities during the West African Ebola outbreak.
This state of the art facility will drive innovation and evidence-based solutions for a growing challenge, by attracting external researchers and enabling them to trial approaches that would be impractical and disruptive to assess in a working ward environment. We expect the facility to be operational towards the end of 2020, based initially at PHE Porton. The project team will work actively with the research community to identify research questions to improve understanding of transmission and potential interventions that could help reduce the numbers of HCAIs and will include locating the facility alongside NHS Trusts where there are particular HCAI challenges or outbreaks.
Developing and evaluating novel therapies
Active engagement with the research community is also essential to identify and evaluate novel therapeutic approaches, such as those aiming to work by altering the microbiome, through the action of host defence peptides or in conjunction with other components of the human immune system. PHE will provide access to researchers wanting to evaluate such strategies on up-to-date clinical isolates of antibiotic-resistant bacteria, in models for difficult to treat infections, such as biofilms.
Accessing such strains and models can be difficult for researchers, especially those who are new to the field and this will allow a range of novel approaches to be explored. This open-innovation approach will stimulate research in this area and provide high quality data to support future development of successful strategies.
We need to take bold steps to reduce antibiotic over-use and preserve them for when we really need them. Through this innovative new approach, we seek to build on the huge progress that has been made so far in understanding the impact of prescribing on resistance and in the development and evaluation of new interventions and begin benefitting the lives of patients sooner.

1 comment
Comment by Guillermo Chussir posted on
I never thought that big data could be useful in the fight against antibiotic resistance. That's awesome!